Proteome activity landscapes of tumor cell lines determine drug responses
2020-07-20
Martin Frejno, Chen Meng, Benjamin Ruprecht, Thomas Oellerich, Sebastian Scheich , Karin Kleigrewe , Enken Drecoll, Patroklos Samaras, Alexander Hogrebe, Dominic Helm, Julia Mergner, Jana Zecha, Stephanie Heinzlmeir, Mathias Wilhelm, Julia Dorn, Hans-Michael Kvasnicka, Hubert Serve, Wilko Weichert, Bernhard Kuster
Nat Commun., 11, 3636, 2020
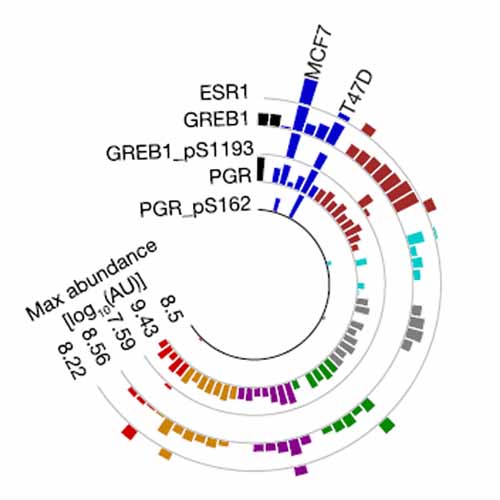
Integrated analysis of genomes, transcriptomes, proteomes and drug responses of cancer cell lines (CCLs) is an emerging approach to uncover molecular mechanisms of drug action. We extend this paradigm to measuring proteome activity landscapes by acquiring and integrating quantitative data for 10,000 proteins and 55,000 phosphorylation sites (p-sites) from 125 CCLs. These data are used to contextualize proteins and p-sites and predict drug sensitivity. For example, we find that Progesterone Receptor (PGR) phosphorylation is associated with sensitivity to drugs modulating estrogen signaling such as Raloxifene. We also demonstrate that Adenylate kinase isoenzyme 1 (AK1) inactivates antimetabolites like Cytarabine. Consequently, high AK1 levels correlate with poor survival of Cytarabine-treated acute myeloid leukemia patients, qualifying AK1 as a patient stratification marker and possibly as a drug target. We provide an interactive web application termed ATLANTiC (http://atlantic.proteomics.wzw.tum.de), which enables the community to explore the thousands of novel functional associations generated by this work.